Overview
Comparative Developmental Biology is an intensive two-week laboratory course for graduate students in year two of their studies or beyond and post-docs, who seek a broad training in experimental approaches to developmental questions across diverse study organisms.
Course Date: October 4 - 18, 2026
Application due date: July 26, 2026
Course description:
This intensive comparative developmental biology course is designed for graduate students in their second or later year of Ph.D studies and post-docs. The two weeklong course will provide exposure to a combination of well-established and emerging developmental systems. Students will develop advanced experimental embryology skills—many of which are transferable across organisms—in the handling and cellular/genetic manipulation of embryos, including microinjection, lineage tracing, microdissection, cell transplantation, in situ hybridization, CRISPR/Cas mutagenesis, and 3D in vivo imaging. Students will develop an enhanced appreciation of the advantages each species offers, will be trained to think more comparatively (in a phylogenetic context), and will gain an appreciation of how best to select the appropriate species to address a specific question. They will be exposed to classic, recent, and developing methodologies and techniques and will learn about exciting ongoing research using these approaches. Developing and completing a short independent or team-based research project will enhance skills in hypothesis generation and experimental design.
Financial Information
Cost: $4000.00 including room and board
Please Note: Financial Aid is not offered for this course
Course Objectives
During this two-week course, students will gain hands-on familiarity with a range of traditional and emerging model systems in developmental biology. They will gain experience in microscopy, image acquisition and analysis, as well as in embryological, pharmacological, and molecular genetic approaches to address key questions in development and regeneration. Students will understand the specific advantages of each model system, gain insights into the types of problem that are currently being researched in each model, and gain experience with key cross-cutting techniques such as CRISPR/Cas mutagenesis, lineage tracing, and in situ hybridization. Students will also enhance their hypothesis building and experimental design and analysis skills through weeklong independent or small-team projects.
The following models are expected to be available in 2024: Zebrafish, Drosophila, Parhyale hawaiensis, Butterfly, Nematostella, Squid, and Skate.
Course Directors:


Nipam Patel
MBL Director
My lab works to uncover both developmental and evolutionary mechanisms in a variety of animals that help us understand the generation of biodiversity, with a recent focus on the crustacean, Parhyale, and several species of butterflies. We use Parhyale to understand the role of Hox genes and other developmental genes in building and evolving body plans and also investigate the mechanisms that allows this species to regenerate its germline. With butterflies, we are working to discover how single scale cells create the impressive nanostructures they use for structural coloration, without which they would be greatly restricted in the pallet of colors available to them.


Vicky Prince
Professor, Organismal Biology & Anatomy, University of Chicago
At the Prince lab we take a cellular, molecular, and comparative approach to the study of developmental processes. Our research program uses rapidly developing, transparent—and very beautiful—embryos of the zebrafish, Danio rerio. Zebrafish embryos are ideal for high resolution live imaging approaches, and these techniques are helping us to understand dynamic cellular processes that build complex structures during embryonic development—especially of the nervous system and cranial structures. Our imaging experiments are complemented by the use of increasingly powerful molecular genetic and transgenic tools, such as CRISPR/Cas technology, which allow us to interrogate the molecular basis of developmental processes. Please take a look around our pages to learn more about our ongoing studies.
Course Faculty:


Karen Echeverri
Associate Scientist, Bell Center
The ability to regenerate complex tissue has fascinated scientists for a long time. The Echeverri lab is interested in the cell and molecular mechanisms driving regeneration. We study regeneration in axolotls; salamanders well known for their ability to functionally regenerate multiple body parts, including limbs, tail, heart, eyes and jaw and in addition can repair lesions in the brain and heal all wounds without forming scar tissue. My group uses cell and molecular tools combined with in vivo imaging and genomics approaches to decipher the key circuitry and cellular mechanisms that are essential to promote functional regeneration.


Amy Herbert
Assistant Professor of Organismal Biology and Anatomy, University of Chicago
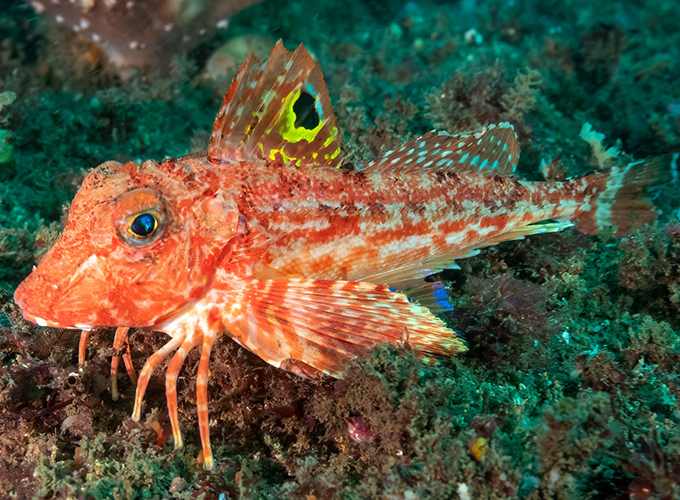
In the lab of evolutionary genetics, we investigate the molecular and genetic mechanisms that drive the evolution of novel vertebrate traits using a combination of model and non‑model organisms. A recently developed research organism in the lab is the sea robin, a marine fish that exhibits multiple evolutionary innovations, including leg‑like structures that allow the fish to walk and taste food on the ocean floor. Using sea robins alongside stickleback fish, zebrafish, and other unique species, we address several fundamental questions in evolutionary biology, including whether similar evolutionary outcomes arise by the same molecular routes, whether new traits are controlled by many genes or by a few key loci, which classes of genes or elements and which types of mutations drive evolutionary change, and how gene regulatory networks evolve to produce novel traits.

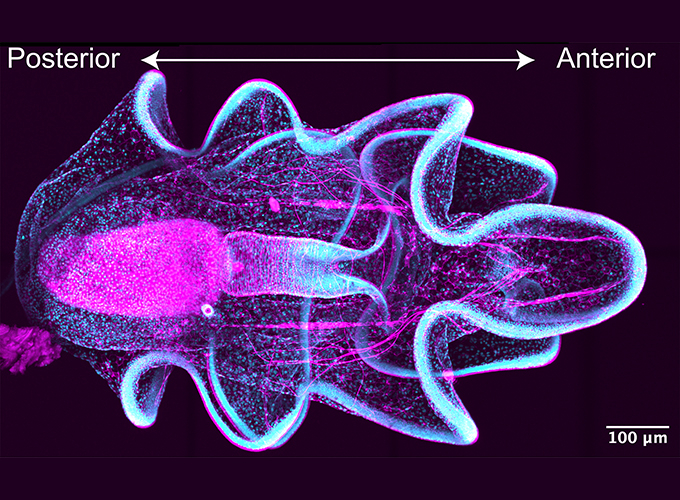
Margherita Perillo
Research Scientist, Bell Center
Epithelial tubes are the building blocks that make virtually any organ across animals, such as lungs, heart, kidneys and more. The Perillo Lab investigates the conserved mechanisms that regulate proper tube morphogenesis in the embryo and maintain healthy organs in the adult. We leverage the larva of sea star Patiria miniata, marine invertebrates that are optically clear, have a simple tubular organ that is easy to access, and are biomedically relevant as sea stars are evolutionarily close to vertebrates. Our current projects combine molecular and cellular tools with imaging and transcriptomic approaches to understand: 1) how cell lineages change from a simple tube to a fully differentiated organ, 2) the mechanisms that keep organ cells in an epithelial state, and 3) how the sea star larva develops a specialized organ for environmental sensing that is vital for survival.
